To discover the potential capabilities and scientific significances of peroxisomes throughout lung most cancers growth and development, we investigated the expressional profiles of peroxisome pathway genes and their correlations with scientific options in non-small cell lung most cancers (NSCLC).
The RNA-seq knowledge of NSCLC together with lung squamous carcinoma (LUSC) and lung adenocarcinoma (LUAD) sufferers with their scientific data had been downloaded from The Cancer Genome Atlas (TCGA). Gene expression comparisons between tumor and regular samples had been carried out with edgeR bundle in R software program and the outcomes of the 83 peroxisome pathway genes had been extracted.
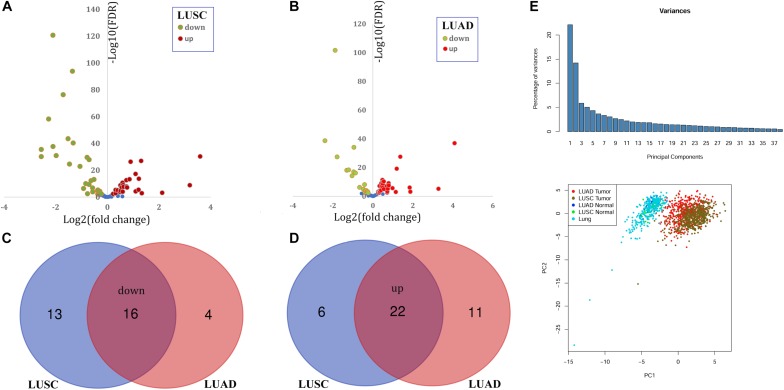
Through Venn diagram evaluation, 38 widespread differentially expressed peroxisome pathway genes (C-DEPGs) in NSCLC had been recognized. Principal parts evaluation (PCA) was carried out and the 38 C-DEPGs might discriminate NSCLC tumors from the non-tumor controls nicely.
Through Kaplan-Meier survival and Cox regression analyses, 11 of the C-DEPGs had been proven to have prognostic results on NSCLC general survival (OS) and had been thought of as key C-DEPGs (Okay-DEPGs). Through Oncomine, Human Protein Atlas (HPA) and the Clinical Proteomic Tumor Analysis Consortium (CPTAC), three Okay-DEPGs (HSD17B4, ACAA1, and PXMP4) had been confirmed to be down-regulated in NSCLC at each mRNA and protein stage.
Their dy-regulation mechanisms had been revealed by way of their correlations with their copy quantity variations and methylation standing. Their potential capabilities in NSCLC had been explored by way of their NSCLC-specific co-expression community evaluation, their correlations with immune infiltrations, immunomodulator gene expressions, MKI67 expression and their associations with anti-cancer drug sensitivity. Our findings advised that HSD17B4, ACAA1, and PXMP4 could be new markers for NSCLC analysis and prognosis and may present new clues for NSCLC therapy.
Quantitative proteomics reveals distinct molecular signatures of various cerebellum-dependent studying paradigms.
The cerebellum improves motor efficiency by adjusting motor achieve appropriately. As de novo protein synthesis is important for the formation and retention of recollections, we hypothesized that motor studying in the other way would induce a definite sample of protein expression in the cerebellum.
We performed quantitative proteomic profiling to match the extent of protein expression in the cerebellum at 1 and 24 hours after coaching from mice that underwent totally different paradigms of cerebellum-dependent oculomotor studying from particular directional adjustments in motor achieve.
We quantified a complete of 43 proteins that had been considerably regulated in every of the three studying paradigms in the cerebellum at 1 and 24 hrs after studying. In addition, purposeful enrichment evaluation recognized protein teams that had been differentially enriched or depleted in the cerebellum at 24 hrs after the three oculomotor studying, suggesting that distinct organic pathways could also be engaged in the formation of three oculomotor reminiscence.
Weighted correlation community evaluation found teams of proteins considerably correlated with oculomotor reminiscence. Finally, 4 proteins (Snca, Sncb, Cttn, and Stmn1) from the protein group correlated with the educational quantity after oculomotor coaching had been validated by Western blot.
This research supplies a complete and unbiased record of proteins associated to a few cerebellum-dependent motor studying paradigms, suggesting the distinct nature of protein expression in the cerebellum for every studying paradigm.